Immune engineering
Programming cells to cure disease
Welcome to The Century of Biology! This newsletter explores data, companies, and ideas from the frontier of biology. You can subscribe for free to have the next post delivered to your inbox:
After a slight lag from the usual cadence because of a move and a trip (more soon), let’s dive back in with an exploration of immune engineering.
Enjoy! 🧬
The immune system is the Macintosh of human biology. It has been elegantly constructed—by Evolution, of course—to maintain an interface between our bodies and the outer world that is always working to press in. Form is connected to function. Complex tasks are carried out with surprising simplicity. Beyond being essential for our survival, it is a testament to the ingenuity of Nature. Now, it is a major frontier for the future of biotechnology and medicine.
Let’s start this story with T cells—a specialist category of immune cells that are capable of adapting to new molecular threats. This adaptation happens when they are presented with antigens, which are strings of molecules—primarily peptides—that can be recognized as foreign. In a major conceptual leap at the time, several physicians and scientists started to hypothesize that T cells could be programmed to target cancer antigens.
They were right. It is possible to hot-wire a patient’s T cells by removing them from their body, engineering them to express a novel Chimeric Antigen Receptor (CAR), and reintroducing the engineered cells to circulate throughout the blood detecting cancer cells. The CARs are a literal chimera of immune signaling components, with a portion of an antibody engineered to recognize cancer antigens strapped on to parts of the signaling domain of a typical T cell receptor.

After decades of grueling research when the pioneers of this field were treated as heretics and pariahs, we started to see incredible results. Some of the earliest patients dosed with CAR-T therapy are still in remission over a decade later. Looking at the initial trial success, the pioneering physician-scientist Carl June spelled out what this meant: “we can now conclude that CAR T cells can actually cure patients with leukemia.”
It’s worth pausing here to let that sink in. If we have a true science-fiction level therapy in the clinic right now, this is it. We are talking about programming immune cells using genetic engineering to cure a complex and systemic genetic disease—cancer. The thing is, this is only the beginning.
As Ground Truths succinctly summarized, there is a lot of activity in the CAR-T space right now. We’re in the early stages of translating the success in blood cancers (leukemia) into solid tumors and a variety of other diseases, and exploring new ways to develop off-the-shelf cell therapies for manufacturing and scaling purposes. Here, I want to specifically drill down on what the future looks like for immune engineering.
I spend a lot of time in this newsletter talking about the impact of Sequencing, Synthesis, Scale, and Software on the world of biotechnology. The 4-S model represents an emerging integrated tool stack for biological engineering. We can design new components using DNA synthesis and rapidly measure the results of parallelized experiments using DNA sequencing. This introduces a new scale of experimentation that generates enough data for software 1.0 and 2.0 (machine learning) to be highly effective. I’ve become convinced that immune engineering is one of the most promising frontiers for the application of the 4-S stack.
Before the 4-S series, I wrote an essay about a really promising study using ML to learn the grammar of CAR receptor combinations.1 This model produced new CAR-T designs based on an underlying model of how the different components of a signal domain work together. The approach used all of the constituent elements of the 4-S stack, but the full vision of where this all might go only clicked into place after I listened to a recent talk by my friend Daniel Goodman.
Daniel earned his PhD working in the lab of George Church at Harvard. George Church is an absolute giant in the world of genomics. He has been on the frontier of DNA reading, writing, and editing for decades. If there is a lab that embodies the 4-S stack, it is the Church Lab.2 You can see the impact of Daniel’s time in the Church Lab in the way that he describes his work:
Our understanding of how genomes control biological function is limited by our ability to make modifications and test hypotheses. As it becomes cheaper and faster to both chemically synthesize DNA and to read from cells with next-generation sequencing, we can identify relationships between DNA sequences and cellular outcomes at an unprecedented scale. This large amount of high-dimensional data can be combined with advancements in artificial intelligence to rapidly predict phenotypes and generate new sequences to test.
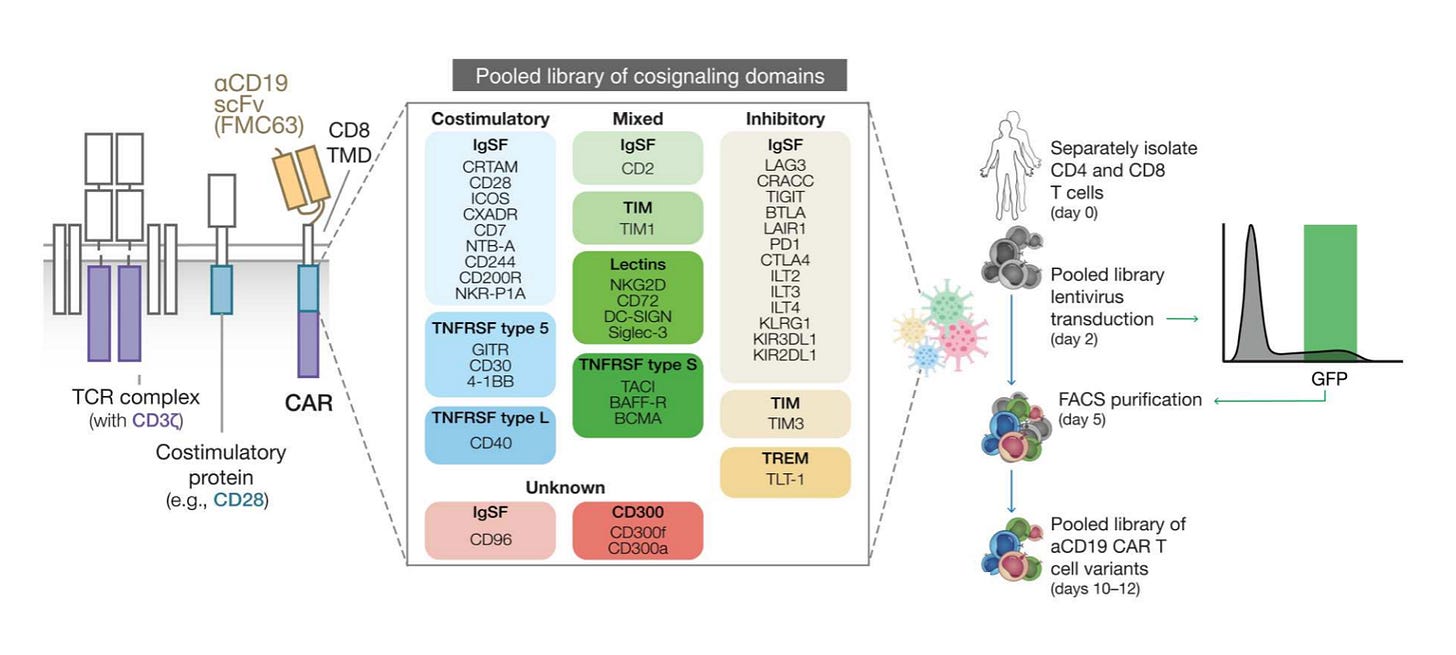
And what is Daniel engineering now? T cells. In recent work, Daniel led an effort to establish an approach for CAR Pooling (nice).3 The main idea is that CARs have been a major success, but still face substantial challenges. There is still high variance in outcomes, and a range of serious patient side effects. One major focus is to improve the design of the costimulatory domains, which are an essential part of the T cell activation process in addition to antigen binding. In this study, they used large-scale DNA synthesis to test a very large number of domains in one big pooled experiment.
This specific technology makes the relationship between genomics and CAR-T therapy very clear. On one hand, we have incredible progress in DNA tech. On the other hand, we have a medical breakthrough that is in serious need of further refinement—specifically in the form of improved genetic engineering. There is a clear convergence. Part of the beauty of this intersection is that CAR-T therapy is a massively high impact application that is capable of being tackled with our current read/write/edit tools. It is an ideal Minimum Viable Product (MVP).
The subsequent work that I wrote up on learning CAR-T grammar is a natural extension into totally synthetic costimulatory domains for even further improvements using AI models. Very cool work from the Chen Lab came out last month in Nature Biotechnology demonstrating massively parallelized gene editing of T cells. This is all to say that large-scale genomics will likely play an important role in developing the next generation of CAR-T therapies.
Let’s zoom out a bit. One thing we can effectively bet on is that we will be able to synthesize longer pieces of DNA more cheaply over time. It’s safe to say gene editing and sequencing will continue to get better too. If CAR-Ts are the MVP for cell engineering, what might come next? For starters, there are a lot of other immune cells to engineer.
It doesn’t require too much imagination to think that other types of immune cells will be engineered into viable therapies. In fact, it’s already happening. Natural killer cell therapies are an active focus of research and development. Why not other cells? What about macrophages? Yep. The list goes on. Fundamentally, we are in the earliest innings of harnessing the immune system for therapeutic purposes.
What about combinations of cells working together to cure more complex diseases? This is clearly a big step up in complexity, but is also an active area of research. One example is the newly launched Hernandez-Lopez Lab at Stanford that is researching “biological circuits in multicellular systems and their applications in biotechnology.”
This increase in complexity between generations of technology is actually a really important detail. One of the views of the a16z Bio investment team is that “the modular aspect of these medicines mean that new applications will be easier to build, reusing and repurposing common components like Lego blocks: once we learn how to deliver a gene to a specific cell in a given disease, it’s significantly more likely we will be able to deliver a different gene to a different cell for another disease.”
It’s possible to develop increasingly sophisticated small molecule drugs, but they can’t be composed together like genetic circuits. In other words, they aren’t Lego blocks—each is a bespoke standalone tool. If this view is true, we should see compounding progress between generations of cell and immune therapies. Different circuits and receptors should be mixed and matched in new ways to cure new diseases.
This is why the intersection of the 4-S stack and immune engineering is so promising. Our DNA technologies are exponentially improving. Immune engineering is a tremendously high-value and high-impact application of genetic engineering and synthetic biology. We should get another level of compounding progress as reusable genetic circuits and components are developed over time.4
It was initially a complete shock to most researchers that the immune system could be hot-wired to treat cancer. Now, the focus has expanded for immune therapy to be a consensus approach for cancer and autoimmune diseases. Will that be the final scope of what is possible? Probably not. Just last year, a study delivered mRNA molecules that programmed T cells still inside the body to express CARs and treat scarring of the heart. If this isn’t a demonstration of compounding progress by combining multiple biotech modalities together, I’m not sure what is.
What if we could engineer every immune cell—and every possible combination of immune cells—to cure every disease? This is clearly an incredibly far out vision for the future of immune engineering, but biology needs more big visions if we are going to reach our full potential.
Thanks for reading this essay about the awesome potential of immune engineering. As always, thanks to my wonderful editor Kelda. If you don’t want to miss upcoming essays, you should consider subscribing for free to have them delivered to your inbox:
Until next time! 🧬
The preprint I covered on January 28, 2022 was ultimately published in Science nearly a year later on December 8, 2022. The lead author of this study, Kyle Daniels, is now a faculty member in the Stanford Genetics Department. You should consider checking out the Daniels Lab which is now hiring, and subscribing to COB to hear about breakthrough science when it first happens!
I’ve been reflecting on how scientific lineage impacts how one approaches problems. A lot of my initial to exposure to the world of 4-S came during my time in the UW Genome Sciences Department. There, seeing Jay Shendure’s work (also trained with Church), building tools with Brian Beliveau (from the adjacent Wu and Yin labs), and collaborating on projects with these groups gave me a heavy dose of the 4-S philosophy—and showed me the generality of DNA-based technology.
I’ve linked to the preprint for open access. This work was published in Science Translational Medicine with some additional data, but the preprint still accurately describes the technology.
This is partly inspired by Jack Clark’s mental model of AI progress. When things start to feel crazy fast, it’s probably something that is compounding. If we do our jobs right, programmable medicines should behave this way too.




May I ask for a piece of advice? What books (scientific, or other sources) would you recommend to read to establish some foundation about biology, DNA and so on for someone who has next to zero previous knowledge in biology but studied physics and has experience in AI and big data? Thank you
Thank you ! Happy to have discovered your newsletter.